Which targets can AMPIVIEW™ probes detect?
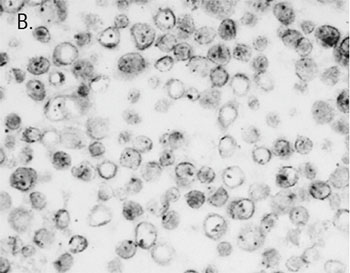
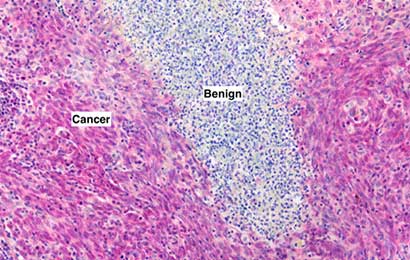
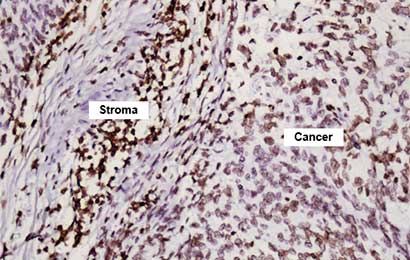
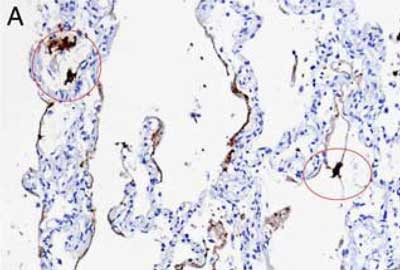
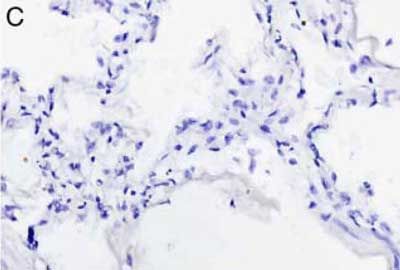
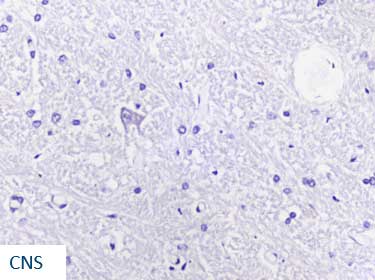
Our AMPIVIEW™ probes based on Enzo’s proprietary LoopRNA™ ISH technology offer outstanding, sensitivity for in situ hybridization, allowing chromogenic detection of nucleic acid targets with a sensitivity that is on par with standard PCR methods – but on tissue slides with all the benefits provided by spatial biology. AMPIVIEW™ probes can detect both DNA and RNA targets, regardless of whether these are genomic targets or expressed copies of RNA, including but not limited to genomic sequences, mRNA, cytosolic viral genomes, mitochondrial DNA.
|
|
How sensitive are AMPIVIEW™ probes based on LoopRNA™ ISH technology?
AMPIVIEW™ probes display extraordinarily high sensitivity, allowing successful chromogenic detection and a strong, easy to visualize staining on par with the sensitivity of standard PCR.
|
|
How are AMPIVIEW™ probes labeled?
The outstanding sensitivity of our AMPIVEW™ probes for in situ hybridization is rooted in Enzo’s proprietary LoopRNA™ ISH technology, which utilizes labeled loops with a high spatial density of label molecules, which are intermittently located throughout the specific hybridization sequence of each probe. These labeled loops can be labeled with either biotin or digoxigenin. This LoopRNA™ ISH technology results on sensitivity on par with standard PCR methods.
|
|
What is the ideal probe length for AMPIVIEW™ probes?
Sense-probes for genomic targets ideally utilize about 3kb of hybridization sequence, while antisense expression probes, which target DNA and RNA, utilize 2kb of hybridization sequence or less, depending on the level of expression of the respective mRNA.
|
|
How are AMPIVEW™ probes detected?
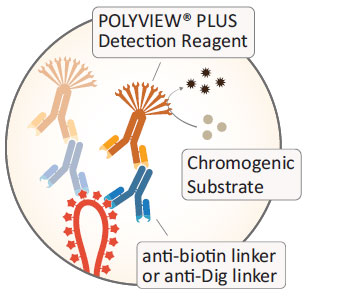
Our AMPIVIEW™ probes are labeled with either biotin or digoxigenin. For biotin-labeled probes, we developed both streptavidin-based detection with our SAVIEW® PLUS detection reagents as well as immunogenic detection with anti-biotin linker antibodies and our POLYVIEW® PLUS detection solutions. Digoxigenin-detection is achieved with anti-digoxigenin linker antibodies in combination with our POLYVIEW® PLUS detection solutions. Regardless of the utilized detection reagent, our HIGHDEF® chromogens offer crisp color chromogenic staining in a broad palette of colors. Of course, AMPIVIEW™ probes can be detected with a broad variety of third-party detection solutions for biotin or digoxigenin labels, including fluorescent detection.
|
|
Which tissue preparation does AMPIVIEW™ require?
Our AMPIVIEW™ probes are typically optimized for formalin-fixed, paraffin-embedded tissue samples and suited for a wide variety of tissue types. Additionally, they are perfectly suited for cryopreserved tissue sections as well, although we do not typically validate this type of tissue specimen in-house. Due to the relative standard and very flexible protocol for in situ hybridization with our AMPIVIEW™ probes, they are theoretically adaptable to a wide variety of cell and tissue specimen and histological techniques, regardless of species origin or fixation and embedding method, as long as the nucleic acids in the sample are preserved.
|
|
What is the volume required for hybridization?
We typically provide our AMPIVIEW™ probes as 2-mL ready-to-use solutions for 20-40 tests. A volume of 50-100 µL per slide has shown to be sufficient for a broad variety of applications and liquid handling procedures for manual and automated workflows, regardless of whether liquid is contained by a hydrophobic barrier, under a cover (such as glass cover slips), or protected from evaporation by an oily film. Depending on the utilized procedure, size of the tissue sections and other variables, significantly smaller volumes might be sufficient.
|
|
Does AMPIVIEW™ require proprietary reagents?
Our biotin- or digoxigenin-labeled AMPIVIEW™ probes can be detected with a broad variety of detection solutions designed to detect the respective labels. AMPIVIEW™-driven chromogenic in situ hybridization provides outstanding results with our SAVIEW® PLUS and POLYVIEW® PLUS detection solutions in combination with our HIGHDEF® chromogens for crisp, strong, and colorful staining that eliminate data ambiguity. However, AMPIVIEW™ probes can be combined with any ISH or IHC solution suitable for biotin or digoxigenin, including fluorescent detection.
|
|
Is AMPIVIEW™ compatible with autostainer instrumentation?
The AMPIVIEW™ workflow allows a high degree of flexibility regarding the respective workflow and the degree of automation. All our AMPIVEW™ products are suitable for manual workflows and a variety of open automated staining instruments. Depending on the autostainer platform, our technical support might provide useful help for the optimization up to established protocols. However, validation of any given workflow is required to maximize results.
|
|
Do you provide positive or negative control probes?
Depending on the target and tissue of choice, we can provide negative and positive control probes. For general purposes, we recommend to use a universal negative scrambled control probe and one of our catalogue probes of ubiquitously expressed targets, such as GAPDH or ubiquitin.
|
|
Is AMPIVIEW™ in situ hybridization influenced by single point mutations?
No. Due to the nature of the AMPIVIEW™ probes and the length of the hybridization area, single point mutations do not influence the signal obtained with Enzo’s AMPIVIEW™ probes. This makes our AMPIVIEW™ probes a great tool to analyze gene targets and expression irrelevant of minor variations, and for example allows the simultaneous detection of virus variants.
|
|
Can AMPIVIEW™ probes be used to detect genomic translocation?
Dual probes utilizing biotin and digoxigenin labels may potentially be used to detect translocation, if the spatial separation between both probe locations is large enough. However, procedures will require careful validation by the end user. Break point localization will be approximate due to the nature of the AMPIVIEW™ probes.
|
|
What is the Quality Control (QC) process for AMPIVIEW™ probes?
The purity of our AMPIVIEW™ probes is determined by A260/280 spectroscopy and gel electrophoresis. Performance is analyzed by dot blot hybridization tests. In situ hybridization on cell or tissue is optional.
|
|








 What is in situ hybridization and what are the recent advancements with this indispensable technique?
What is in situ hybridization and what are the recent advancements with this indispensable technique?