Polymerase chain reaction (PCR) is a relatively simple and widely used molecular biology technique to amplify and detect DNA and RNA sequences. Compared to traditional methods of DNA cloning and amplification, which can often take days, PCR requires only a few hours. PCR is highly sensitive and requires minimal template for detection and amplification of specific sequences. Basic PCR methods have further advanced from simple DNA and RNA detection. Below, we have provided an overview of the different PCR methods and the reagents we provide at Enzo Life Sciences for your research needs. We aim to help scientists quickly access PCR reagents to use in their next research project!
PCR
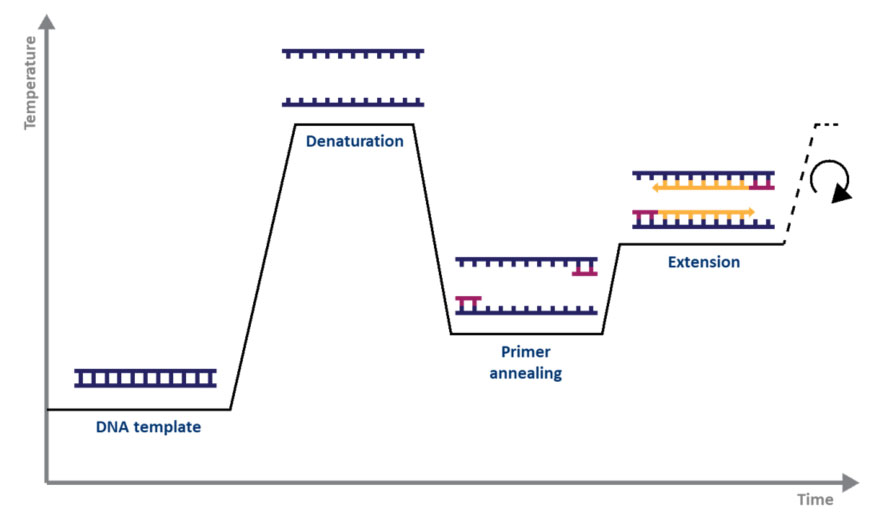
For standard PCR, all you need is a DNA polymerase, magnesium, nucleotides, primers, the DNA template to be amplified, and a thermocycler. The PCR mechanism is as simple as its purpose: 1) double-stranded DNA (dsDNA) is heat denatured, 2) primers align to the single DNA strands, and 3) the primers are extended by DNA polymerase, resulting in two copies of the original DNA strand. The denaturation, annealing, and elongation process over a series of temperatures and times is known as one cycle of amplification (Fig. 1).
|
Figure 1. Schematic representation of a cycle of amplification by PCR.
|
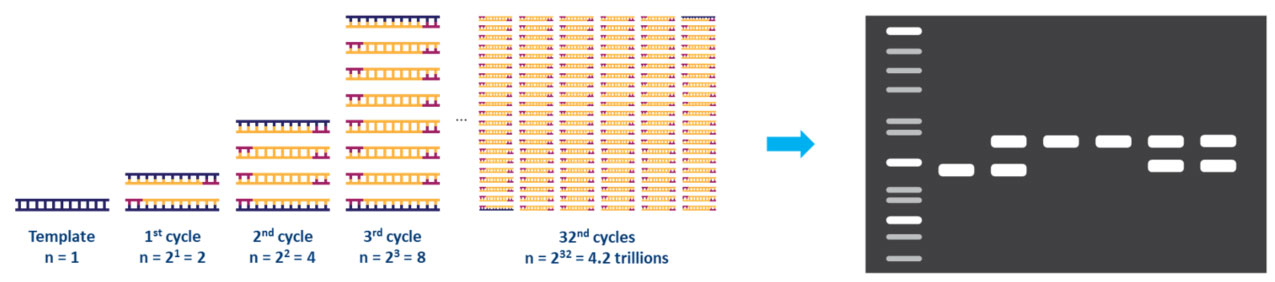
Each step of the cycle should be optimized for the template and primer set used. This cycle is repeated approximately 20-40 times, and the amplified product can then be analyzed, typically by agarose gel (Fig. 2).
|
Figure 2. Amplification of a DNA template by PCR and analysis by agarose gel electrophoresis.
|
As PCR is a highly sensitive method and very small volumes are required for single reactions, the preparation of a master mix for several reactions is recommended. The master mix must be well mixed and then split by the number of reactions, ensuring that each reaction will contain the same amount of enzyme, dNTPs, and primers. Many suppliers, such as Enzo Life Sciences, also offer PCR mixes that already contain everything except primers and the DNA template.
Guanine/Cytosine-rich (GC-rich) regions represent a challenge in standard PCR techniques. GC-rich sequences are more stable than sequences with lower GC content. Furthermore, GC-rich sequences tend to form secondary structures, such as hairpin loops. As a result, GC-rich double strands are difficult to completely separate during the denaturation phase. Consequently, DNA polymerase cannot synthesize the new strand without hindrance. A higher denaturation temperature can improve this, and adjustments towards a higher annealing temperature and shorter annealing time can prevent the unspecific binding of GC-rich primers. Additional reagents can enhance the amplification of GC-rich sequences. DMSO, glycerol, and betaine help to disrupt the secondary structures that are caused by GC interactions and thereby facilitate the separation of the double strands.
Hot Start PCR
Unspecific amplification is a problem that can occur during PCR. Most DNA polymerases that are used for PCR work best at temperatures around 68°C to 72°C. The enzyme can, however, also be active at lower temperatures, although to a lower degree. At temperatures far below the annealing temperature, primers can bind non-specifically and lead to non-specific amplification, even if the reaction is set up on ice. This can be prevented by using polymerase inhibitors that dissociate from the DNA polymerase only once a certain temperature is reached, hence the term hot start PCR. The inhibitor can be an antibody that binds the polymerase and denatures at the initial denaturation temperature (95°C typically).
High Fidelity Polymerase
While DNA polymerases amplify fairly accurately to the original template sequence, mistakes in nucleotide matching can occur. Mismatches in applications such as cloning can result in truncated transcripts, and mistranslated or inactive proteins downstream. To avoid these mismatches, polymerases with a “proofreading” activity have been identified and incorporated into the workflow. The first proofreading polymerase, Pfu, was identified in 1991 in Pyrococcus furiosus. This Pfu enzyme has a 3’ to 5’ exonuclease activity. As the DNA is amplified, the exonuclease removes mismatched nucleotides at the 3’ end of the strand. The correct nucleotide is then replaced, and DNA synthesis continues. The identification of incorrect nucleotide sequences is based on the binding affinity for the correct nucleoside triphosphate with the enzyme, where inefficient binding slows the synthesis and allows for the correct replacement. The proofreading activity of Pfu polymerase results in fewer errors in the final sequence compared to Taq DNA polymerase. In recent years, other proofreading enzymes have been identified, and modifications of the original Pfu enzyme have been made to further reduce the error rate during DNA amplification.
RT-PCR
Reverse transcription PCR, or RT-PCR, allows the use of RNA as a template. An additional step allows the detection and amplification of RNA. The RNA is reverse transcribed into complementary DNA (cDNA), using reverse transcriptase. The quality and purity of the RNA template are essential for the success of RT-PCR. The first step of RT-PCR is the synthesis of a DNA/RNA hybrid. Reverse transcriptase also has an RNase H function, which degrades the RNA portion of the hybrid. The single-stranded DNA molecule is then completed by the DNA-dependent DNA polymerase activity of the reverse transcriptase into cDNA. The efficiency of the first-strand reaction can affect the amplification process. From here on, the standard PCR procedure is used to amplify the cDNA. The possibility to revert RNA into cDNA by RT-PCR has many advantages, and it is primarily used for gene expression analysis. RNA is single-stranded and very unstable, which makes it challenging to work with. It commonly serves as a first step in qPCR, which quantifies RNA transcripts in a biological sample.
qPCR and RT-qPCR
Quantitative PCR (qPCR) is used to detect, characterize and quantify nucleic acids for numerous applications. In RT-qPCR, RNA transcripts are often quantified by reverse transcribing them into cDNA first, as described above, and then qPCR is subsequently carried out. As in standard PCR, DNA is amplified by three repeating steps: denaturation, annealing, and elongation. However, in qPCR, fluorescent labeling enables the collection of data as PCR progresses. This technique has many benefits due to the range of methods and chemistries available.
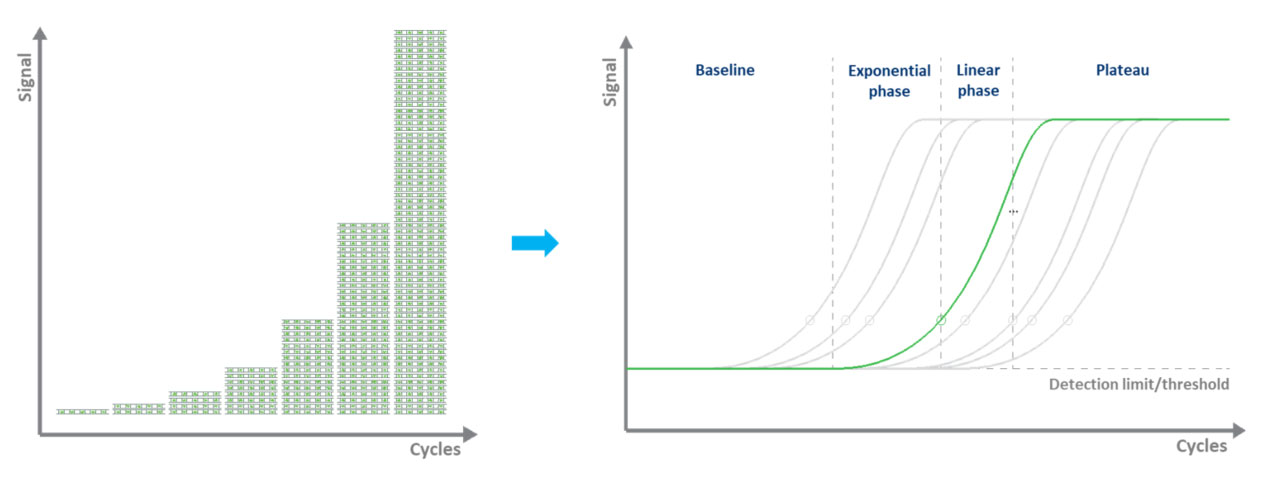
In dye-based qPCR (typically green), fluorescent labeling allows the quantification of the amplified DNA molecules by employing the use of a dsDNA binding dye. During each cycle, the fluorescence is measured. The fluorescence signal increases proportionally to the amount of replicated DNA. Hence, the DNA is quantified in “real-time” (Fig. 3). The disadvantages to dye-based qPCR are that only one target can be examined at a time and that the dye will bind to any ds-DNA present in the sample.
|
Figure 3. Amplifying a DNA template by qPCR and measuring the fluorescence signal in real-time.
|
In probe-based qPCR, many targets can be detected simultaneously in each sample, but this requires optimization and design of a target-specific probe(s) used in addition to primers. Several types of probe designs are available, but the most common type is a hydrolysis probe, which incorporates a fluorophore and quencher. Fluorescence resonance energy transfer (FRET) prevents the emission of the fluorophore via the quencher while the probe is intact. However, during the PCR reaction, the probe is hydrolyzed during primer extension and amplification of the specific sequence it is bound to. The cleavage of the probe separates the fluorophore from the quencher and results in an amplification-dependent increase in fluorescence (Fig. 4). Thus, the fluorescence signal from a probe-based qPCR reaction is proportional to the amount of the probe target sequence present in the sample. Because probe-based qPCR is more specific than dye-based qPCR, it is often the technology used in qPCR-based diagnostic assays.

|
Figure 4. Differences between dye-based and probe-based qPCR.
|
Isothermal Amplification
The PCR mentioned above techniques requires expensive thermocycling equipment to accurately ramp up and down chamber temperatures for the denaturation, annealing, and extension steps. A number of techniques have been developed that do not need such precise devices and can be performed in a simple water bath or even within the cells of interest. These techniques are collectively called isothermal amplification and work based on exponential, linear, or cascade amplification.
The best-known type of isothermal amplification is loop-mediated isothermal amplification, or LAMP. LAMP uses exponential amplification at 65⁰C to amplify template DNA or RNA. When performing LAMP, four to six primers complementary to regions of the target DNA are used with a DNA polymerase to synthesize new DNA. Two of these primers have complimentary sequences that recognize sequences in the other primers and bind them, allowing for a “loop” structure to form in the newly synthesized DNA that then aids primer annealing in subsequent rounds of amplification. LAMP can be visualized by multiple methods, including fluorescence, agarose gel electrophoresis, or colorimetry. The ease of visualizing and detecting the presence or absence of product by colorimetry and the lack of expensive equipment required made LAMP a suitable option for SARS-CoV-2 testing in areas where clinical lab testing was not readily available, or storage and transport of samples was not feasible, or in labs that did not previously have PCR thermocycling equipment.
What is Digital PCR?
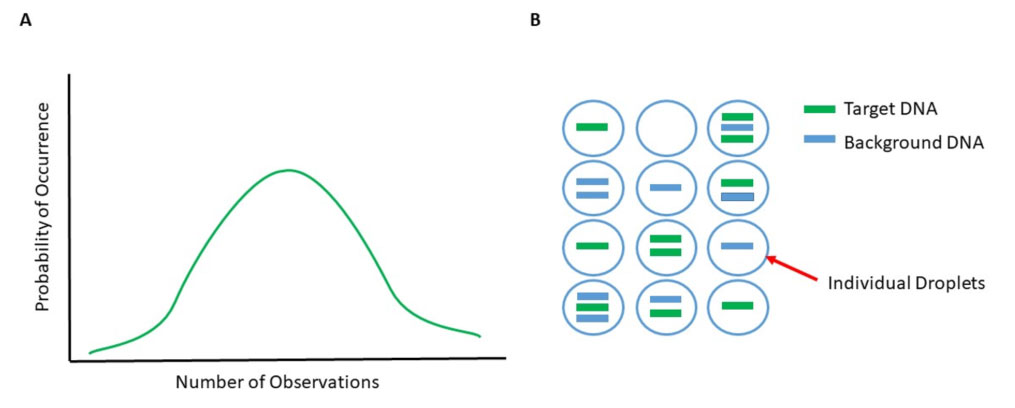
While qPCR and RT-qPCR are considered quantitative, they provide only relative quantitation as the results are reported with respect to the expression of a control, or “housekeeper” gene, or a standard curve produced through serial dilutions. To provide an absolute quantitation, digital PCR was developed. With digital PCR, the total number of initial DNA targets can be quantitated, rather than a calculation based on the amount of PCR product. This quantitation of initial DNA targets was developed by Sykes et al. 1 using sample dilution and Poisson statistics. In Poisson statistics, the probability of something occurring (or, in the case of digital PCR, the expression of the target gene of interest) is calculated based on the occurrence within a known set of events (Fig. 5A). The known set of events was determined by a limited dilution of the initial sample. In 1999, Vogelstein and Kinzler2 took this a step further by diluting the samples enough that the amount of template in each could be amplified by PCR without competition from other targets, increasing the sensitivity of digital PCR. The DNA is recognized by a fluorescent probe specific to the target. These initial samples are now no longer simply diluted, but partitioned into fractions by several different technological methods. Some methods fractionate the sample to be applied on a microarray chip with thousands of probes which can then be amplified by PCR. Others use microfluidics to dissect the sample where the amplification can occur.
One standard commercial method is to disperse the template DNA into thousands of liquid droplets, with each droplet containing as few template molecules as possible (Figure 1B). The presence or absence of target DNA in each droplet would be considered one of the “events” used by Poisson statistics. Each individual droplet has the DNA inside amplified, and the number of droplets that have the target of DNA of interest is determined. Calculations are made based on how many of the total number of droplets contain the target DNA. This process is thus known as droplet-based digital PCR, or ddPCR). Since the quantitation is determined based on the presence or absence of the target, there is less variability from PCR efficiency variables that could affect the cycle threshold (Ct) and, thus, less variation in the calculation.
|
Figure 5. Poisson distribution and partitioning sample DNA. (A) When provided a known number of observations, the probability of a future occurrence can be calculated via Poisson statistics. Increasing the number of observations increases the tendency to a normal distribution. (B) Partitioning of sample DNA into thousands of individual droplets for ddPCR results in minimal amounts of DNA within each droplet for amplification.
|
The advantages of digital PCR have allowed for its use to be incorporated into several applications. The low amounts of RNA found in liquid biopsy samples make digital PCR ideal for these types of diagnostics, the analysis of single-cell samples, and detecting low amounts of pathogens during early infection. Digital PCR also allows for the detection of rare sequences and copy number variations since there is less background DNA in each partition to mask the specific amplification.
Further details can be found on our website:
AMPIGENE® PCR & qPCR Solutions. Enzo’s mission is to provide low-cost, easily adaptable, and effective solutions to the diagnostic market. Our
AMPIPROBE® platform uses a novel PCR design to enable the quantification of nucleic acids for clinically relevant targets. Our assays are easily adaptable for laboratory use and cost-effective, without compromising on quality and performance. Compatible with open PCR platforms, AMPIPROBE® assays can be validated on existing instrumentation, eliminating the need for capital expenditures.
The summary table below highlights the PCR products available for PCR, RT-PCR and qPCR. Please click on the product of interest for more information or contact our
Technical Support Team for further assistance.
Check out our
10 tips for successful Real Time qPCR!













